ReCER Science
Our rapidly changing world requires rapidly evolving science-based solutions.
We use innovative science and technology to investigate the factors impacting the distribution and assembly of plant species.
Our research outcomes inform the restoration and conservation of resilient ecosystems.
We establish technologically advanced large-scale knowledge infrastructure systems that inform the preservation and restoration of plant diversity.
- Restore and Renew
- Conservation genomics for threatened flora
- Evolutionary resilient germplasm collections
- Rainforest evolution and conservation
- Myrtle rust management
- Genome research
- Aboriginal food plant dispersal
- Environmental Microbiome
- Speciation and species delineation
- Weed genomics for improved management
- Phytophthora management
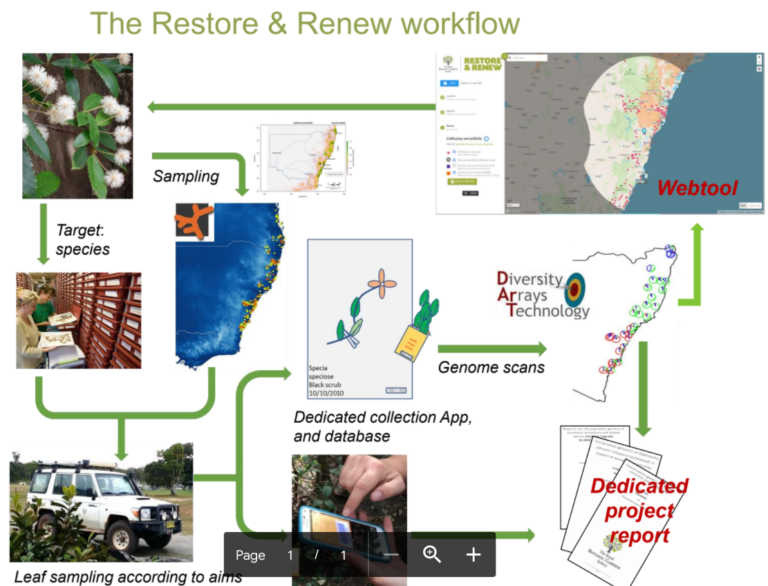
Restore and Renew
Restore & Renew responds to the need for ecological restoration practitioners to incorporate the latest science into their toolkit, helping them to restore diverse, resilient and adaptable ecosystems. Resilient ecosystems need to be made up of species that are not only adapted to the local geology, climate and soil, but to future conditions as well. Restore and Renew acquires empirical knowledge on genetic diversity, habitat availability and distributional patterns across multiple species to deliver restoration guidance to practitioners in easy to use publicly available web tools.
More information:
Conservation genomics for threatened flora
ReCER supports the NSW Government Saving our Species program through the provision of genomic information to help guide the recovery and long term management of threatened flora. Owing to decreasing costs and increased efficiency, it is now conceivable that conservation genomic information can be used to improve the effectiveness of recovery programs for many, if not most, threatened plants. To support biodiversity managers, we developed a simple, standardized workflow for genomic research that guides the efficient collection, analysis and application of genomic information across disparate threatened plants. We argue for a shift away from asking whether genomic information is needed or justified, towards consideration of the questions that need to be addressed.
More information:
View ReCER’s conservation genomics reports to date and a list of taxa currently being worked on.
A conservation genomics workflow to guide practical management actions (Research paper)
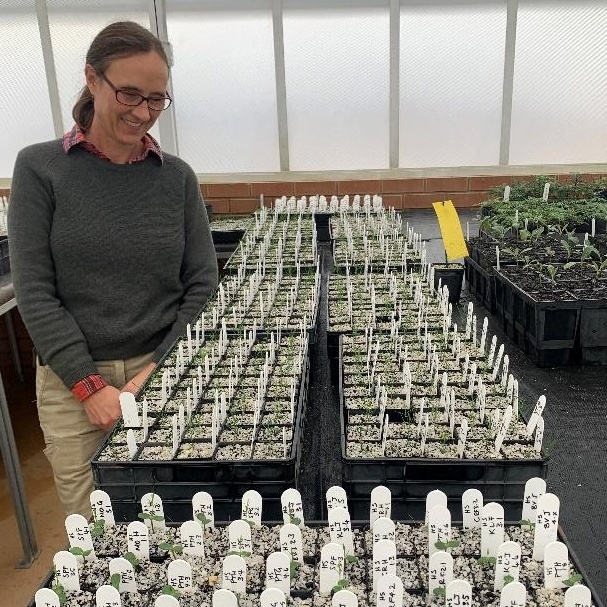


Supporting evolutionary resilient germplasm collections
The long-term success and resilience of ecological restoration and conservation relies heavily on the quality of the material available through germplasm collections. High quality material will survive the initial establishment phase and will be able to adapt to future challenges. Whether stored seed, living collections or seed production areas, germplasm collections that maximise genetic diversity are more likely to facilitate adaptation than collections that are evolutionarily unrepresentative. We are using a combination of genomic and experimental data to deliver collecting strategies that will optimise genetic diversity of germplasm collections.
Rainforest evolution and conservation
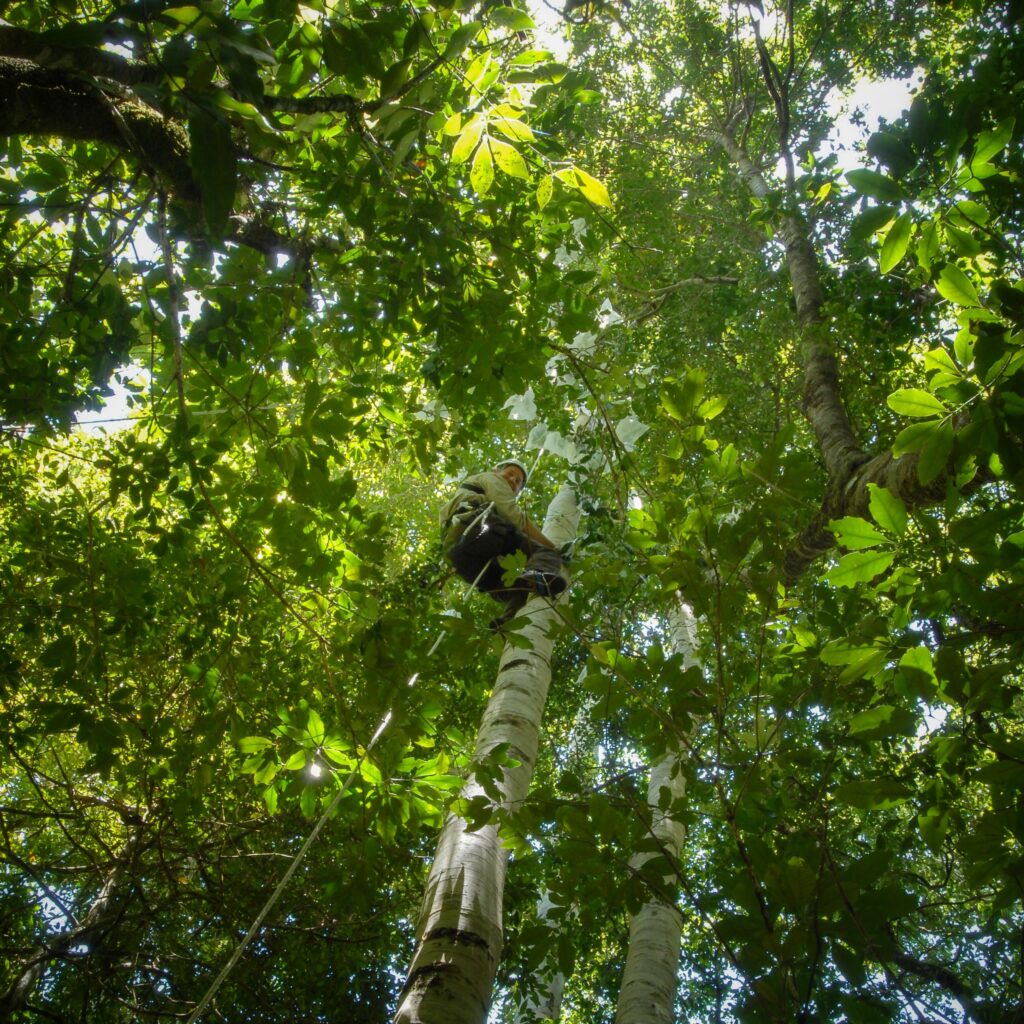
Australian rainforests are highly diverse, with the subtropical forests of northern NSW being particularly rich in Gondwanan lineages. Unfortunately, during the last 150 years, the impacts of logging, clearing, urbanisation and fire have significantly reduced the condition and extent of rainforest vegetation in Australia. Climate change further threatens the integrity and long-term sustainability of these ecosystems. Understanding how rainforest communities are assembled, and why these assemblages change through time can guide rainforest preservation and restoration. In particular, we are interested in the impact of dispersal, historical climatic shifts, associations between genes and environment, floristic exchanges between continental floras, and in identifying which rainforest areas have persisted through time (i.e. which areas are long-term refugia). Identifying refugia and understanding the characteristics that make vegetation persist can help us understand which areas are more vulnerable, and identify priority areas for biodiversity conservation
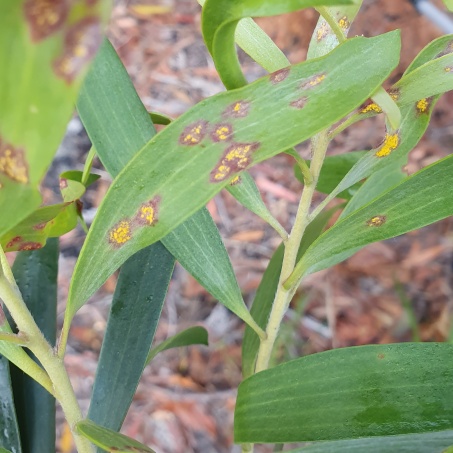


Myrtle rust management
We are helping to conserve plant species that are being impacted by myrtle rust. Myrtle rust is a disease caused by a fungus that was first detected in Australia in 2010. Many Australian species in the myrtle family can be infected, and four species have become critically endangered due to the impacts of the disease. We are partnering with NSW government officers and university researchers to protect genetically diverse collections of these plant species, while exploring options for promoting their resistance to the disease.
Our research is increasingly conducted at whole genome scale, and we have partnered with university researchers to develop genomic resources for a number of our study species. These resources enable studies that aim to identify parts of the genome that influence important outcomes, such as conferring resistance to a disease, or limiting reproduction between different groups of plants (leading to the formation of separate species).
More information:
New discovery gives hope of survival for critically endangered Wollemi Pine’ (Sydney Morning Herald article)
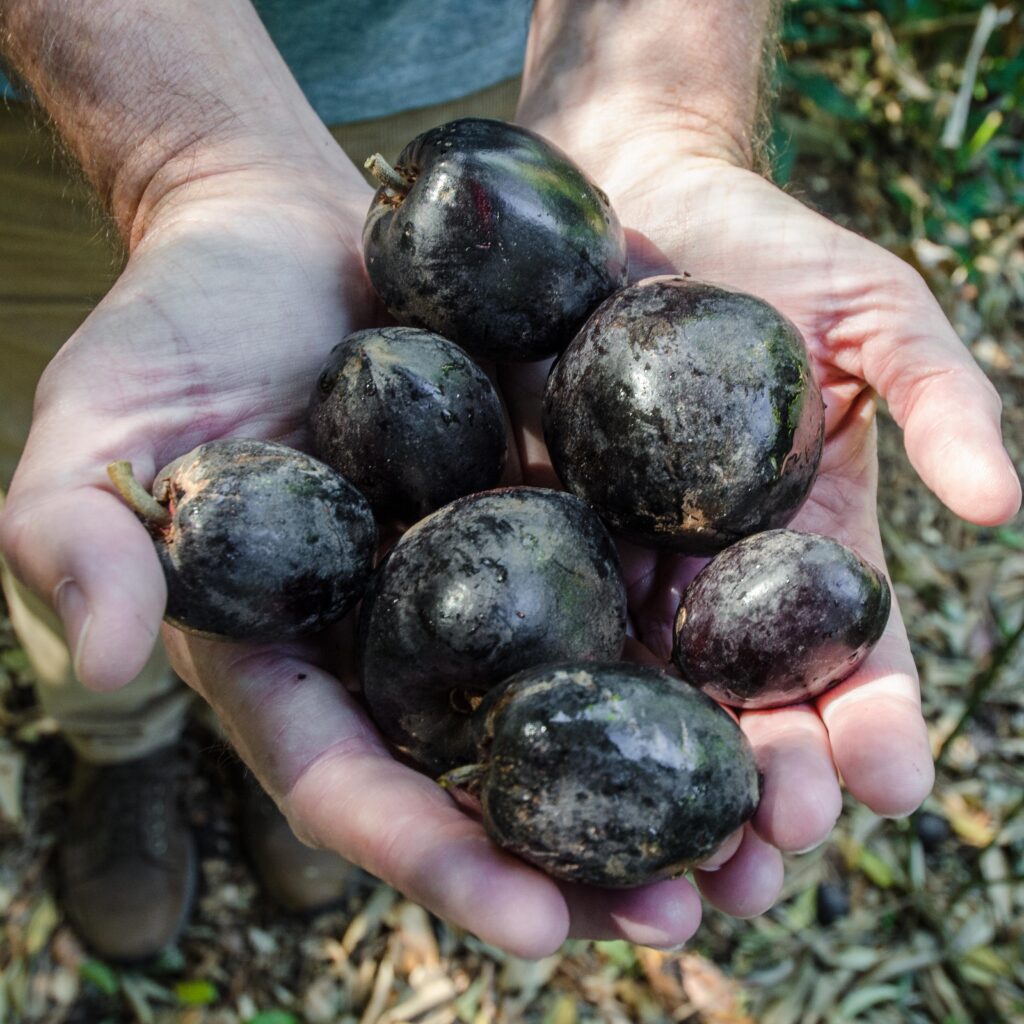


Aboriginal food plant dispersal
This multidisciplinary project aims to retrace the dispersal of native food trees by pre-colonial Aboriginal people, with a focus on Black Bean (Castanospermum australe) and Bunya Pine (Araucaria bidwillii). We are combining plant whole-genome-sequence data with biocultural evidence to investigate whether population genetic links within each species correspond with Aboriginal cultural pathways. We will also estimate the timing of dispersal events with coalescent models to determine whether ancient people expanded the natural range of the two species. The findings from this project will deepen our understanding of ancient plant cultivation practices and how Aboriginal custodianship has influenced the ecology and evolution of the Australian flora.
More information:
The story behind the journey of the black bean tree (Lighthouse article)
Rainforest fruit genetics confirm ancient Aboriginal pathways (watch)
Rainforest fruit genetics reveal ancient Aboriginal pathways (read)
Environmental Microbiome
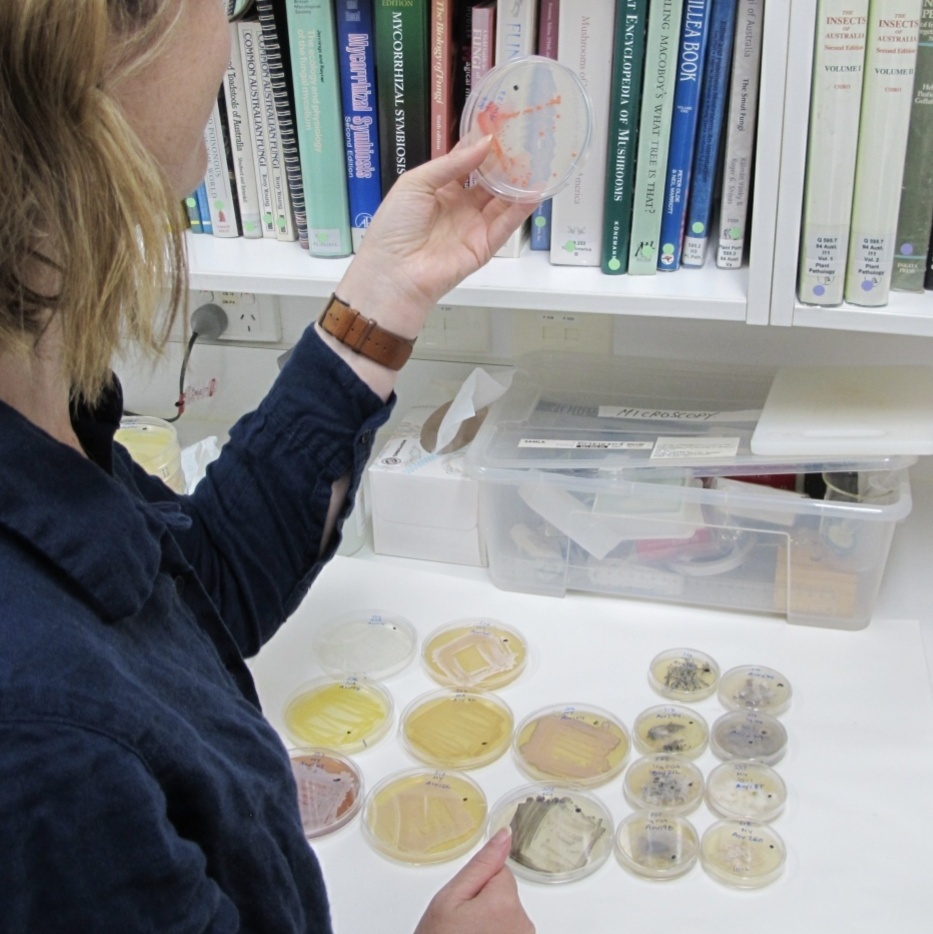
Microbial communities are an essential component of natural ecosystems, associated with increasing ecosystem function and productivity, but are often overlooked in our understanding of ecosystem resilience and conservation. While we are aware of the multitude of roles microbes play in the maintenance of ecosystem processes, little is known about the hidden microbial diversity in Australian native plants and what roles they play. With a focus on endophytes—fungi or bacteria that live within plants—we combine traditional culturing and cutting edge DNA technologies, to increase our understanding of the patterns of microbial diversity, abundance and spatial dynamics in different host tissues across a range of ecosystems. This knowledge provides a more complete understanding of the complexities and diversity in natural ecosystems and how this should be considered in restoration and conservation management.
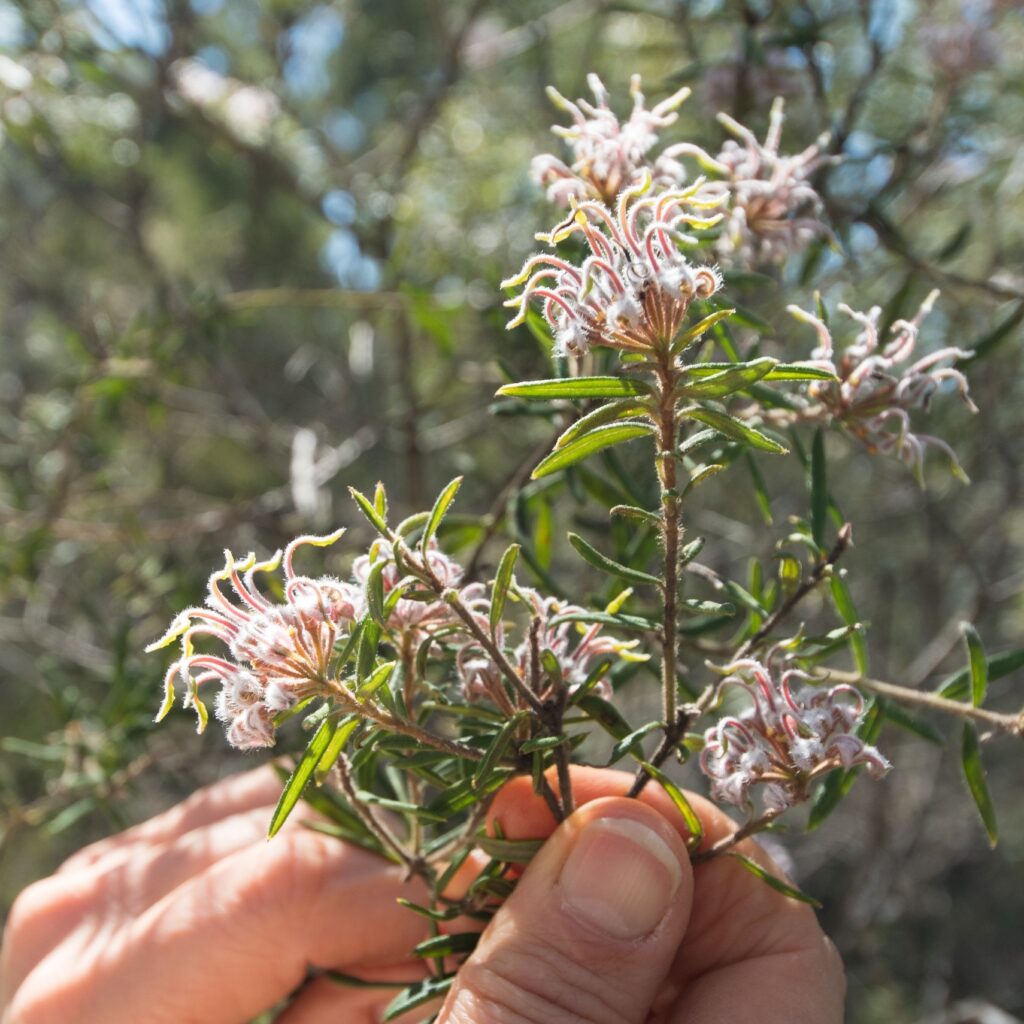


Speciation and species delineation
Defining speciation processes by integrating genomic, morphological and environmental data across a range of native species is critical for understanding evolutionary processes and supporting conservation and management of our unique flora. We explore recent radiations, hybridisation fronts, polyploidisation, adaptive divergences and much more.
More information:
New research into our most iconic tree provides evolutionary insights
Weed genomics for improved management
Invasive plants cost millions of dollars in economic and environmental impacts every year. Understanding a weed’s origin, genetic make-up, and reproductive processes enables more effective management by unlocking targeted control methods (such as natural enemies, i.e. biological control agents). ReCER delivers fundamental insights into the population genomics of invasive plants, which help to advance evidence-based approaches to weed management. Our findings support research partners and stakeholders to develop and adopt innovative solutions to improve control of established weeds.
More information on the project webpage here.
Plus read: Helping scientists control Lantana
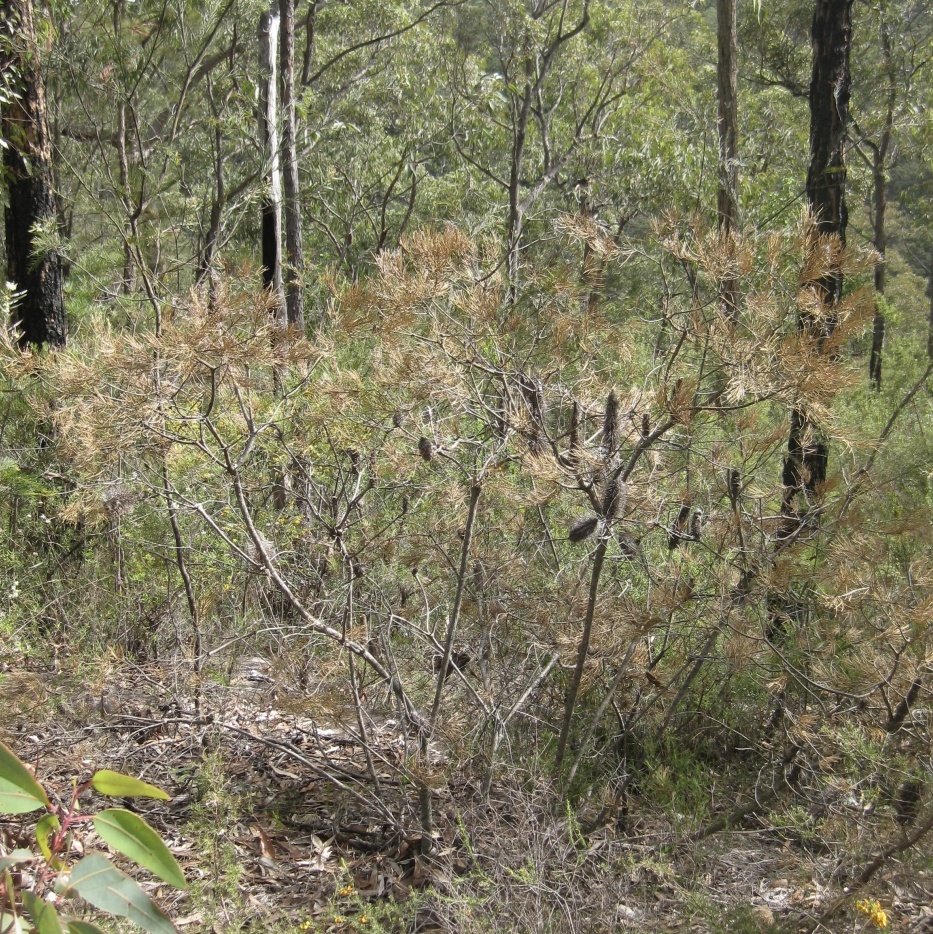


Phytophthora management
Phytophthora is a soil borne plant pathogen that causes devastating root rot and dieback. Phytophthora cinnamomi infects a broad range of plant species (>2000) and occurs widely across Australia. Recognised by both the NSW and Commonwealth Governments as ‘a key threatening process’ to our biodiversity, its impact in natural ecosystems include changes in floristic structure, diversity, and ecological processes, along with changes in animal species abundance and population structure. We are focussing on managing Phytophthora in NSW by further understanding its biology and ecology, determining its susceptibility to threatened species, assessing ways to control its spread and analysing its risks at the landscape scale.
More information:
